ML之RF&XGBoost:基于RF/XGBoost(均+5f-CrVa)算法对Titanic(泰坦尼克号)数据集进行二分类预测(乘客是否生还)
ML之RF&XGBoost:基于RF/XGBoost(均+5f-CrVa)算法对Titanic(泰坦尼克号)数据集进行二分类预测(乘客是否生还)
目录
输出结果






![]()
![]()
![]()





比赛结果

设计思路

核心代码
- rfc = RandomForestClassifier()
- rfc_cross_val_score=cross_val_score(rfc, X_train, y_train, cv=5).mean()
- print('RF:',rfc_cross_val_score)
-
- rfc.fit(X_train,y_train)
- rfc_y_predict = rfc.predict(X_test)
- rfc_submission = pd.DataFrame({'PassengerId': test['PassengerId'], 'Survived': rfc_y_predict})
- rfc_submission.to_csv('data_input/Titanic Data/Titanic_rfc_submission.csv', index=False)
-
-
-
- xgbc = XGBClassifier()
- xgbc_cross_val_score=cross_val_score(xgbc, X_train, y_train, cv=5).mean()
- print('XGBoost:',xgbc_cross_val_score)
-
- xgbc.fit(X_train, y_train)
- xgbc_y_predict = xgbc.predict(X_test)
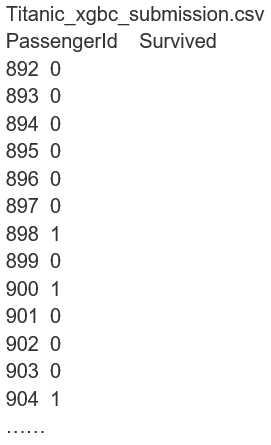
- xgbc_submission = pd.DataFrame({'PassengerId': test['PassengerId'], 'Survived': xgbc_y_predict})
- xgbc_submission.to_csv('data_input/Titanic Data/Titanic_xgbc_submission.csv', index=False)
- class RandomForestClassifier(-title class_ inherited__">ForestClassifier):
- """A random forest classifier.
-
- A random forest is a meta estimator that fits a number of decision tree
- classifiers on various sub-samples of the dataset and use averaging to
- improve the predictive accuracy and control over-fitting.
- The sub-sample size is always the same as the original
- input sample size but the samples are drawn with replacement if
- `bootstrap=True` (default).
-
- Read more in the :ref:`User Guide <forest>`.
-
- Parameters
- ----------
- n_estimators : integer, optional (default=10)
- The number of trees in the forest.
-
- criterion : string, optional (default="gini")
- The function to measure the quality of a split. Supported criteria are
- "gini" for the Gini impurity and "entropy" for the information gain.
- Note: this parameter is tree-specific.
-
- max_features : int, float, string or None, optional (default="auto")
- The number of features to consider when looking for the best split:
-
- - If int, then consider `max_features` features at each split.
- - If float, then `max_features` is a percentage and
- `int(max_features * n_features)` features are considered at each
- split.
- - If "auto", then `max_features=sqrt(n_features)`.
- - If "sqrt", then `max_features=sqrt(n_features)` (same as "auto").
- - If "log2", then `max_features=log2(n_features)`.
- - If None, then `max_features=n_features`.
-
- Note: the search for a split does not stop until at least one
- valid partition of the node samples is found, even if it requires to
- effectively inspect more than ``max_features`` features.
-
- max_depth : integer or None, optional (default=None)
- The maximum depth of the tree. If None, then nodes are expanded
- until
- all leaves are pure or until all leaves contain less than
- min_samples_split samples.
-
- min_samples_split : int, float, optional (default=2)
- The minimum number of samples required to split an internal node:
-
- - If int, then consider `min_samples_split` as the minimum number.
- - If float, then `min_samples_split` is a percentage and
- `ceil(min_samples_split * n_samples)` are the minimum
- number of samples for each split.
-
- .. versionchanged:: 0.18
- Added float values for percentages.
-
- min_samples_leaf : int, float, optional (default=1)
- The minimum number of samples required to be at a leaf node:
-
- - If int, then consider `min_samples_leaf` as the minimum number.
- - If float, then `min_samples_leaf` is a percentage and
- `ceil(min_samples_leaf * n_samples)` are the minimum
- number of samples for each node.
-
- .. versionchanged:: 0.18
- Added float values for percentages.
-
- min_weight_fraction_leaf : float, optional (default=0.)
- The minimum weighted fraction of the sum total of weights (of all
- the input samples) required to be at a leaf node. Samples have
- equal weight when sample_weight is not provided.
-
- max_leaf_nodes : int or None, optional (default=None)
- Grow trees with ``max_leaf_nodes`` in best-first fashion.
- Best nodes are defined as relative reduction in impurity.
- If None then unlimited number of leaf nodes.
-
- min_impurity_split : float,
- Threshold for early stopping in tree growth. A node will split
- if its impurity is above the threshold, otherwise it is a leaf.
-
- .. deprecated:: 0.19
- ``min_impurity_split`` has been deprecated in favor of
- ``min_impurity_decrease`` in 0.19 and will be removed in 0.21.
- Use ``min_impurity_decrease`` instead.
-
- min_impurity_decrease : float, optional (default=0.)
- A node will be split if this split induces a decrease of the impurity
- greater than or equal to this value.
-
- The weighted impurity decrease equation is the following::
-
- N_t / N * (impurity - N_t_R / N_t * right_impurity
- - N_t_L / N_t * left_impurity)
-
- where ``N`` is the total number of samples, ``N_t`` is the number of
- samples at the current node, ``N_t_L`` is the number of samples in the
- left child, and ``N_t_R`` is the number of samples in the right child.
-
- ``N``, ``N_t``, ``N_t_R`` and ``N_t_L`` all refer to the weighted sum,
- if ``sample_weight`` is passed.
-
- .. versionadded:: 0.19
-
- bootstrap : boolean, optional (default=True)
- Whether bootstrap samples are used when building trees.
-
- oob_score : bool (default=False)
- Whether to use out-of-bag samples to estimate
- the generalization accuracy.
-
- n_jobs : integer, optional (default=1)
- The number of jobs to run in parallel for both `fit` and `predict`.
- If -1, then the number of jobs is set to the number of cores.
-
- random_state : int, RandomState instance or None, optional
- (default=None)
- If int, random_state is the seed used by the random number
- generator;
- If RandomState instance, random_state is the random number
- generator;
- If None, the random number generator is the RandomState instance
- used
- by `np.random`.
-
- verbose : int, optional (default=0)
- Controls the verbosity of the tree building process.
-
- warm_start : bool, optional (default=False)
- When set to ``True``, reuse the solution of the previous call to fit
- and add more estimators to the ensemble, otherwise, just fit a whole
- new forest.
-
- class_weight : dict, list of dicts, "balanced",
- "balanced_subsample" or None, optional (default=None)
- Weights associated with classes in the form ``{class_label: weight}``.
- If not given, all classes are supposed to have weight one. For
- multi-output problems, a list of dicts can be provided in the same
- order as the columns of y.
-
- Note that for multioutput (including multilabel) weights should be
- defined for each class of every column in its own dict. For example,
- for four-class multilabel classification weights should be
- [{0: 1, 1: 1}, {0: 1, 1: 5}, {0: 1, 1: 1}, {0: 1, 1: 1}] instead of
- [{1:1}, {2:5}, {3:1}, {4:1}].
-
- The "balanced" mode uses the values of y to automatically adjust
- weights inversely proportional to class frequencies in the input data
- as ``n_samples / (n_classes * np.bincount(y))``
-
- The "balanced_subsample" mode is the same as "balanced" except
- that
- weights are computed based on the bootstrap sample for every tree
- grown.
-
- For multi-output, the weights of each column of y will be multiplied.
-
- Note that these weights will be multiplied with sample_weight
- (passed
- through the fit method) if sample_weight is specified.
-
- Attributes
- ----------
- estimators_ : list of DecisionTreeClassifier
- The collection of fitted sub-estimators.
-
- classes_ : array of shape = [n_classes] or a list of such arrays
- The classes labels (single output problem), or a list of arrays of
- class labels (multi-output problem).
-
- n_classes_ : int or list
- The number of classes (single output problem), or a list containing the
- number of classes for each output (multi-output problem).
-
- n_features_ : int
- The number of features when ``fit`` is performed.
-
- n_outputs_ : int
- The number of outputs when ``fit`` is performed.
-
- feature_importances_ : array of shape = [n_features]
- The feature importances (the higher, the more important the feature).
-
- oob_score_ : float
- Score of the training dataset obtained using an out-of-bag estimate.
-
- oob_decision_function_ : array of shape = [n_samples, n_classes]
- Decision function computed with out-of-bag estimate on the training
- set. If n_estimators is small it might be possible that a data point
- was never left out during the bootstrap. In this case,
- `oob_decision_function_` might contain NaN.
-
- Examples
- --------
- >>> from sklearn.ensemble import RandomForestClassifier
- >>> from sklearn.datasets import make_classification
- >>>
- >>> X, y = make_classification(n_samples=1000, n_features=4,
- ... n_informative=2, n_redundant=0,
- ... random_state=0, shuffle=False)
- >>> clf = RandomForestClassifier(max_depth=2, random_state=0)
- >>> clf.fit(X, y)
- RandomForestClassifier(bootstrap=True, class_weight=None,
- criterion='gini',
- max_depth=2, max_features='auto', max_leaf_nodes=None,
- min_impurity_decrease=0.0, min_impurity_split=None,
- min_samples_leaf=1, min_samples_split=2,
- min_weight_fraction_leaf=0.0, n_estimators=10, n_jobs=1,
- oob_score=False, random_state=0, verbose=0, warm_start=False)
- >>> print(clf.feature_importances_)
- [ 0.17287856 0.80608704 0.01884792 0.00218648]
- >>> print(clf.predict([[0, 0, 0, 0]]))
- [1]
-
- Notes
- -----
- The default values for the parameters controlling the size of the trees
- (e.g. ``max_depth``, ``min_samples_leaf``, etc.) lead to fully grown and
- unpruned trees which can potentially be very large on some data
- sets. To
- reduce memory consumption, the complexity and size of the trees
- should be
- controlled by setting those parameter values.
-
网站声明:如果转载,请联系本站管理员。否则一切后果自行承担。
赞同 0
评论 0 条
- 上周热门
- 如何使用 StarRocks 管理和优化数据湖中的数据? 2672
- 【软件正版化】软件正版化工作要点 2637
- 统信UOS试玩黑神话:悟空 2532
- 信刻光盘安全隔离与信息交换系统 2216
- 镜舟科技与中启乘数科技达成战略合作,共筑数据服务新生态 1092
- grub引导程序无法找到指定设备和分区 743
- WPS City Talk · 校招西安站来了! 15
- 金山办公2024算法挑战赛 | 报名截止日期更新 15
- 看到某国的寻呼机炸了,就问你用某水果手机发抖不? 14
- 有在找工作的IT人吗? 13
- 本周热议
- 我的信创开放社区兼职赚钱历程 40
- 今天你签到了吗? 27
- 信创开放社区邀请他人注册的具体步骤如下 15
- 如何玩转信创开放社区—从小白进阶到专家 15
- 方德桌面操作系统 14
- 我有15积分有什么用? 13
- 用抖音玩法闯信创开放社区——用平台宣传企业产品服务 13
- 如何让你先人一步获得悬赏问题信息?(创作者必看) 12
- 2024中国信创产业发展大会暨中国信息科技创新与应用博览会 9
- 中央国家机关政府采购中心:应当将CPU、操作系统符合安全可靠测评要求纳入采购需求 8
热门标签更多

